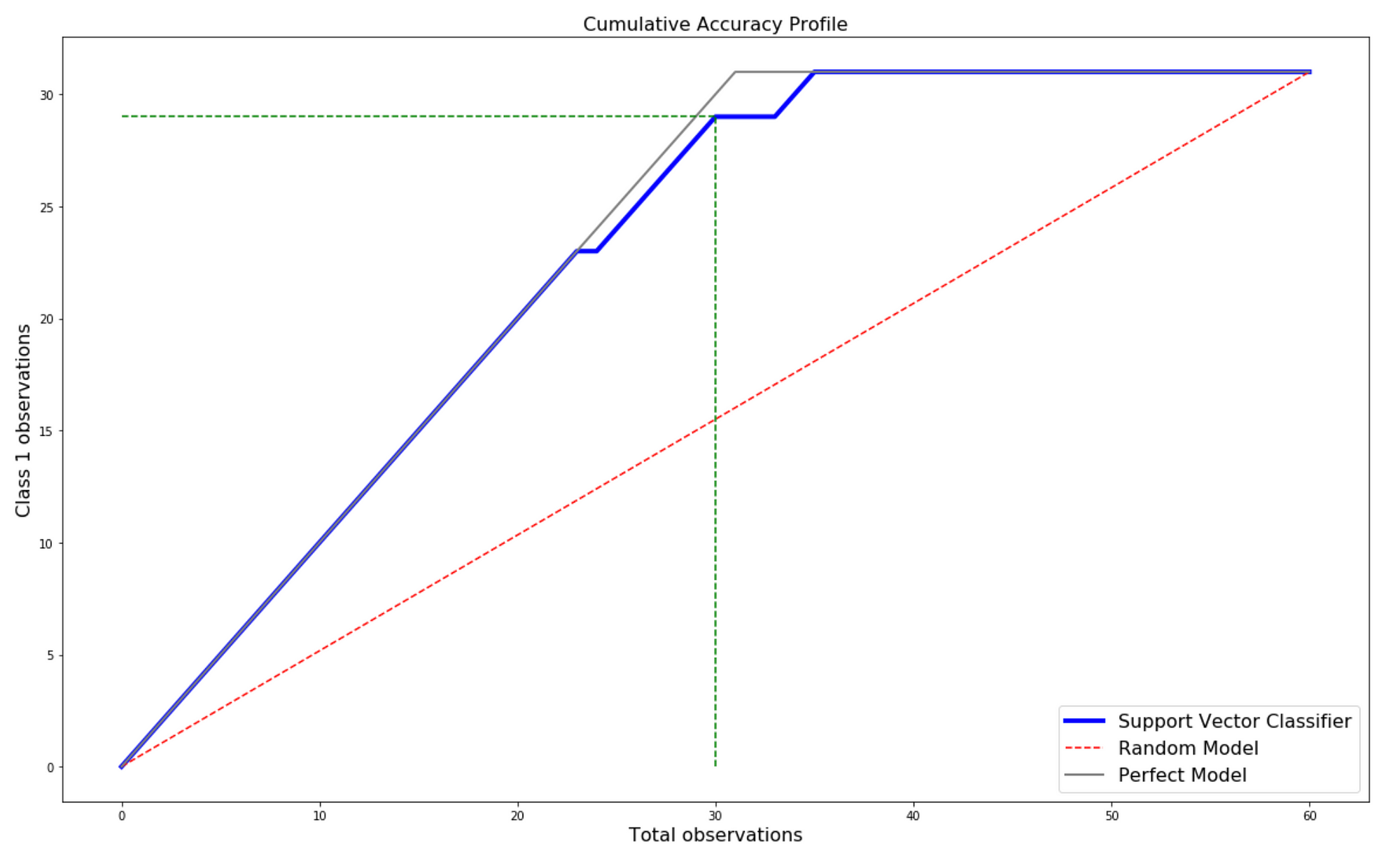
Machine Learning Classifier evaluation using ROC and CAP Curves | by Karan Bhanot | Towards Data Science

RNA on X: "5' cap incorporation is a critical quality attribute in mRNA manufacturing but very little guidance for designing an assay has been available in the literature. This paper tries to

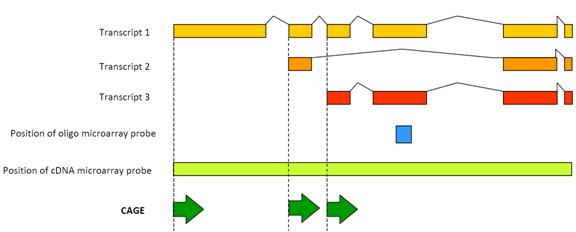
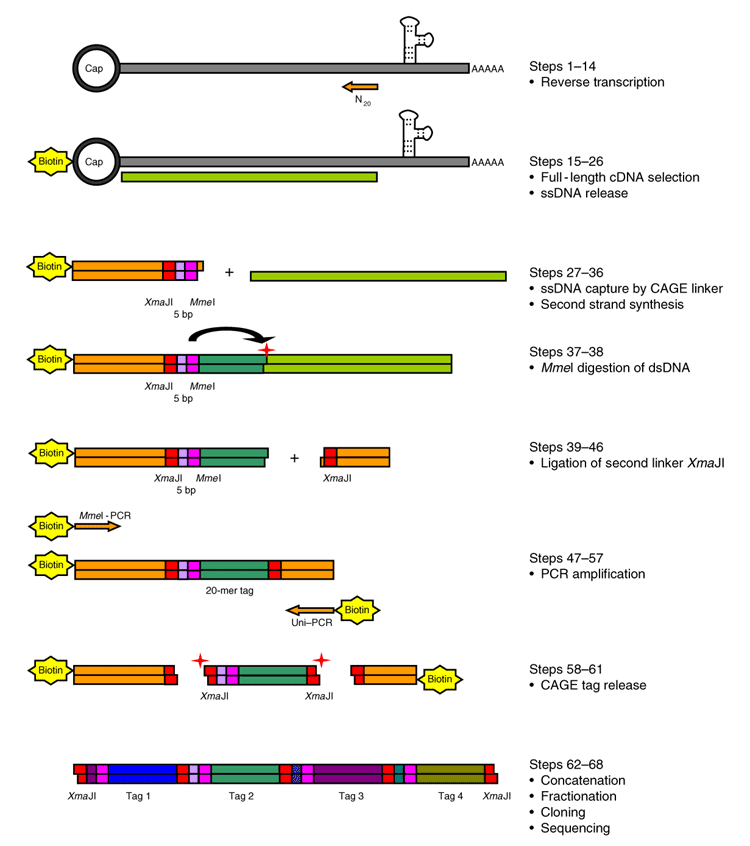
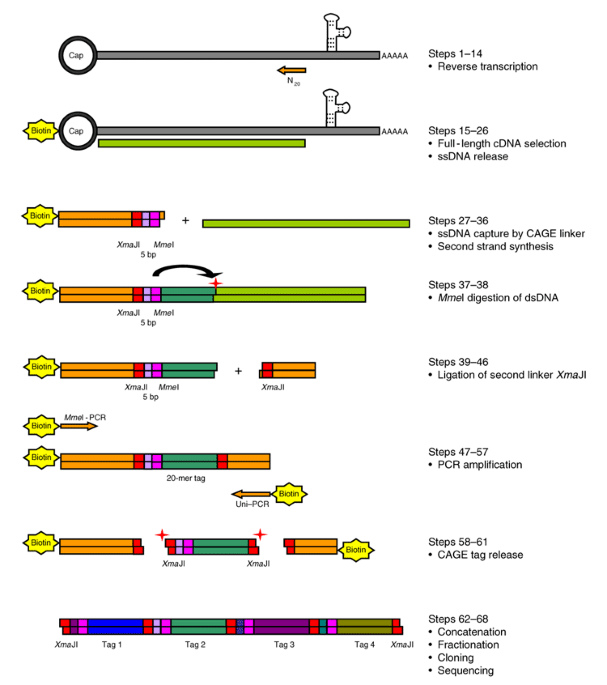
Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage | PNAS

Canonical analysis of principal coordinates (CAP) ordination plot (Bray-Curtis) of fish assemblage data for each experimental treatment at each geographic location.
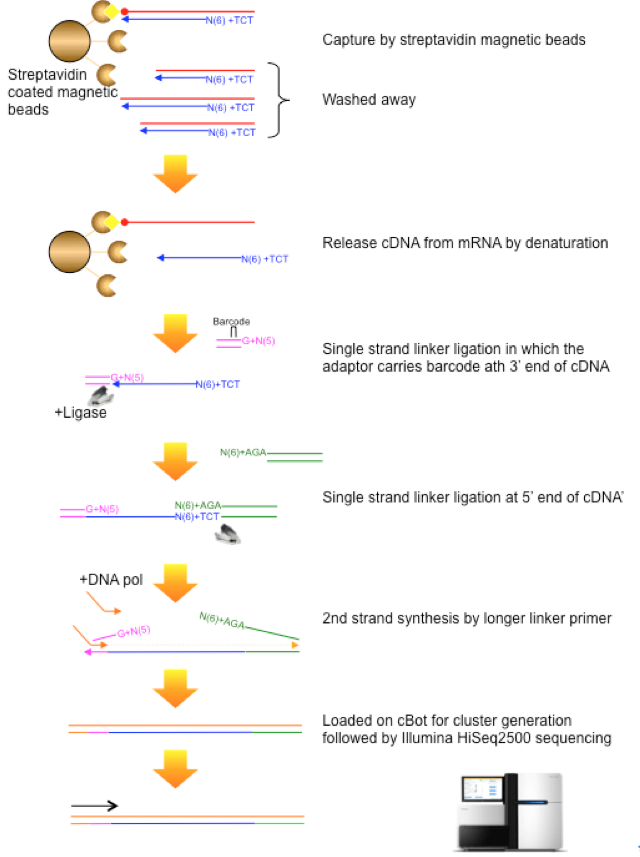
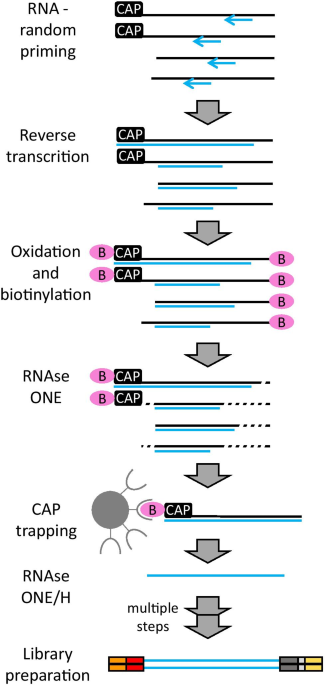
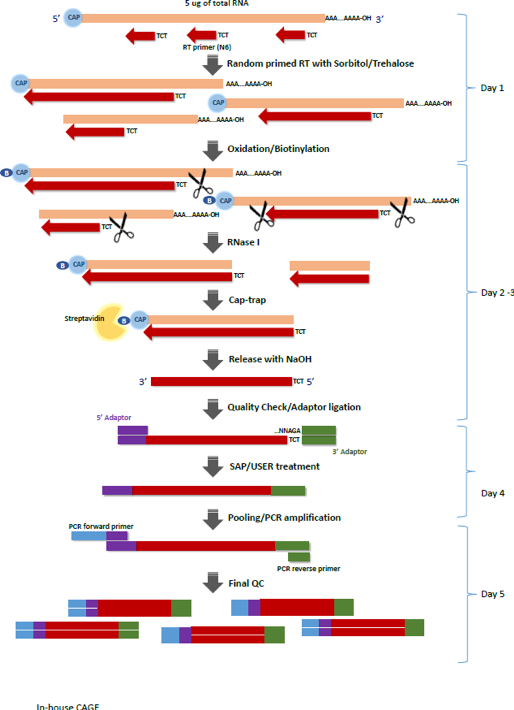
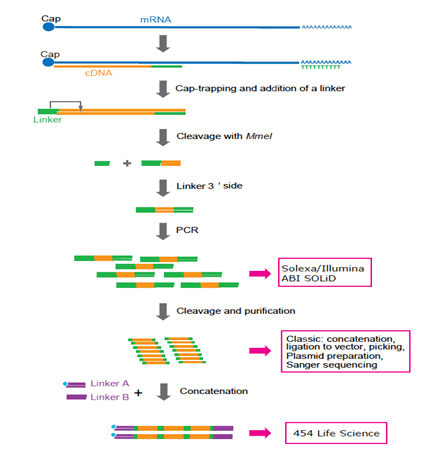
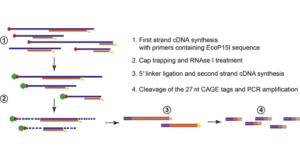
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター

CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology