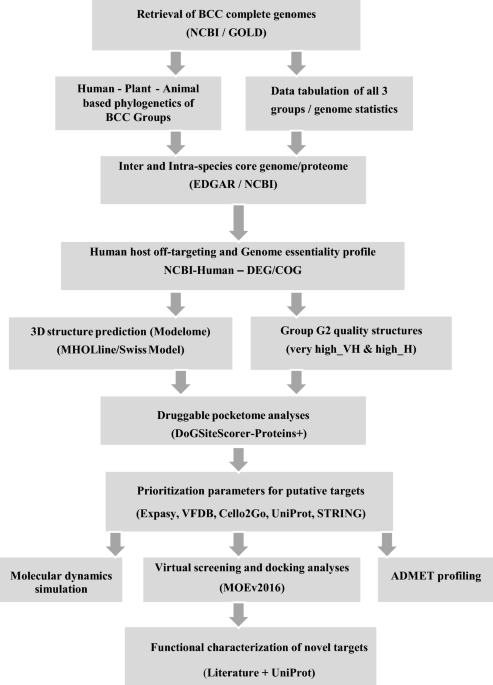
Subtractive sequence analysis aided druggable targets mining in Burkholderia cepacia complex and finding inhibitors through bioinformatics approach | Molecular Diversity
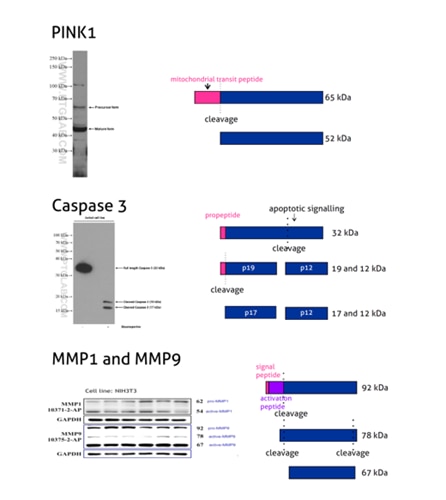
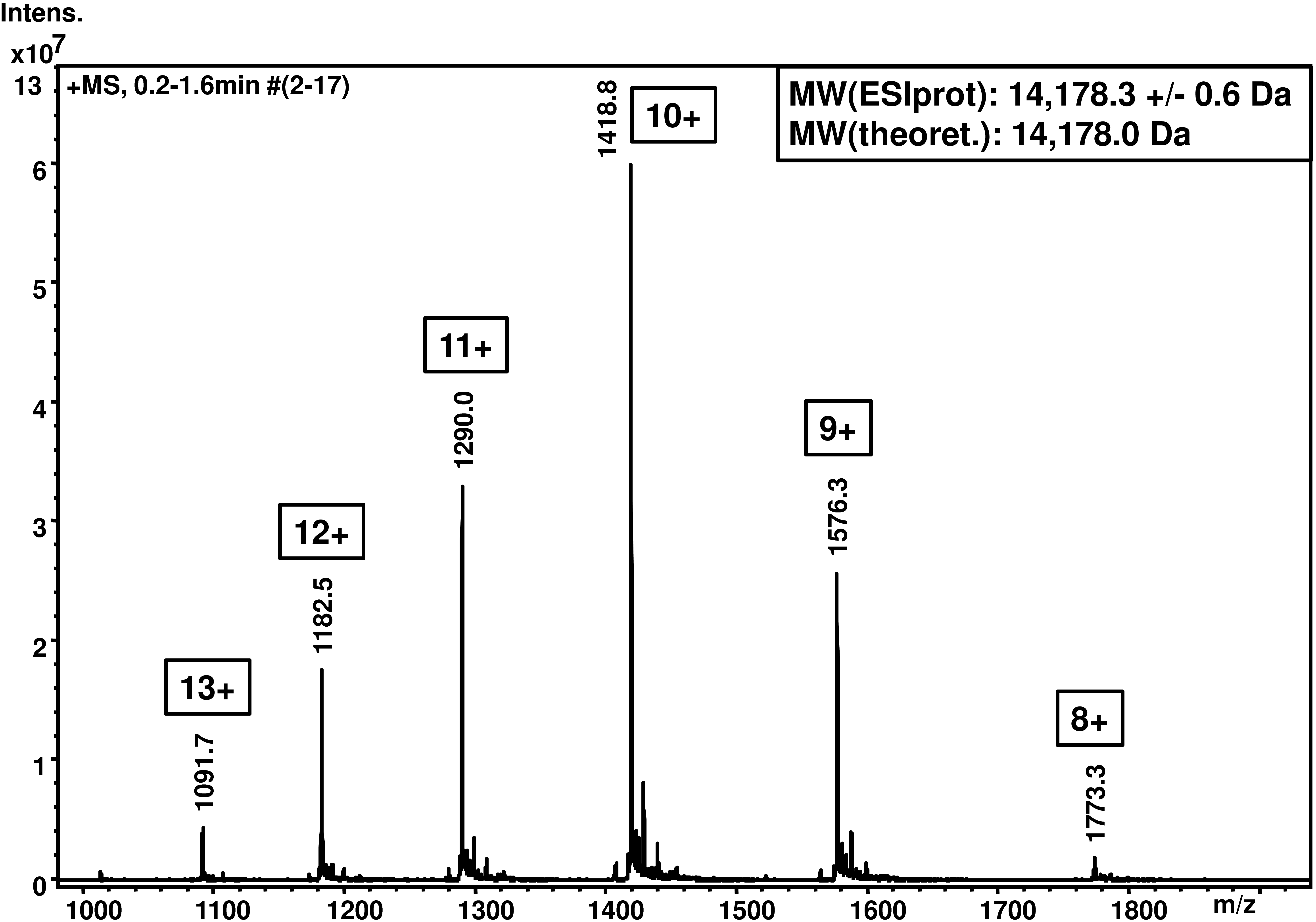
Western Blot Troubleshooting: Why Does The Observed Protein Molecular Weight (MW) Differ From The Calculated One? | Proteintech Group
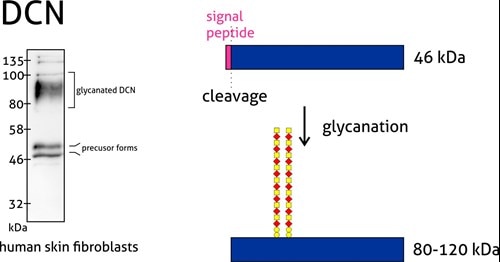
Western Blot Troubleshooting: Why Does The Observed Protein Molecular Weight (MW) Differ From The Calculated One? | Proteintech Group

Why Does the Molecular Weight of My Protein Differ From the Theoretically Expected Weight? | Technology Networks

Expasy ProtParam: pI (isoelectric point), extinction coefficient for UV-based concentration, etc. - YouTube

Bioinformatics Tools and Knowledgebases to Assist Generating Targeted Assays for Plasma Proteomics | SpringerLink