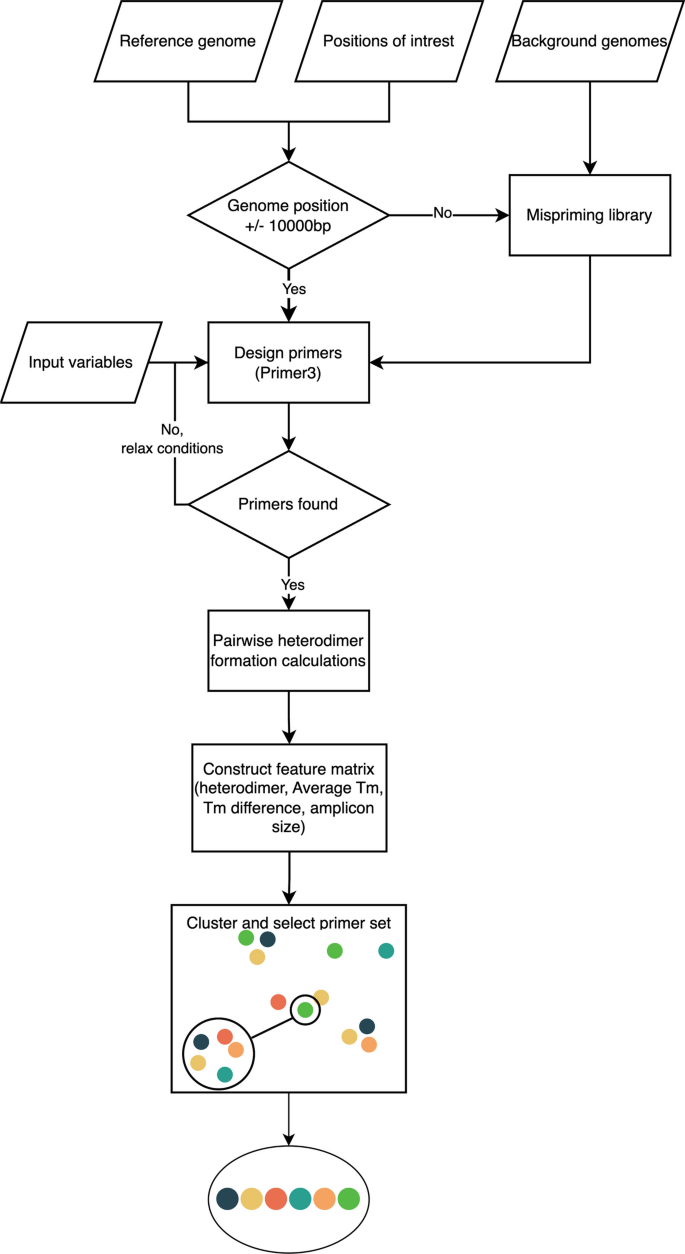
primerJinn: a tool for rationally designing multiplex PCR primer sets for amplicon sequencing and performing in silico PCR | BMC Bioinformatics | Full Text

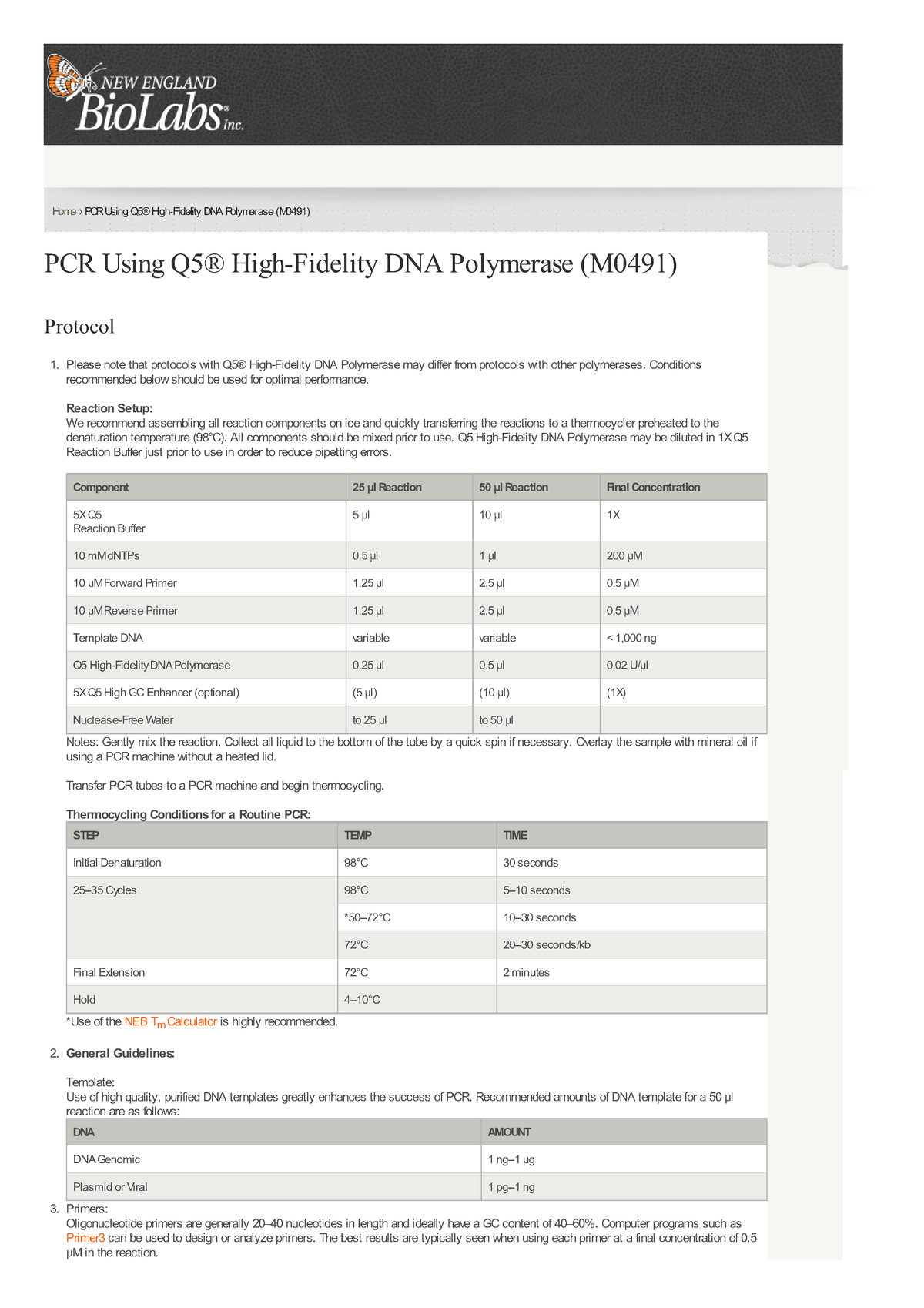
PCR Using Q5U Hot Start High-Fidelity DNA Polymerase (NEB #M0515): Amplification of bisulfite-converted, deaminated, or damaged DNA (Including FFPE DNA)

Having trouble when amplifying GC-rich sequences? Here are four tips on how to troubleshoot – BioNordika
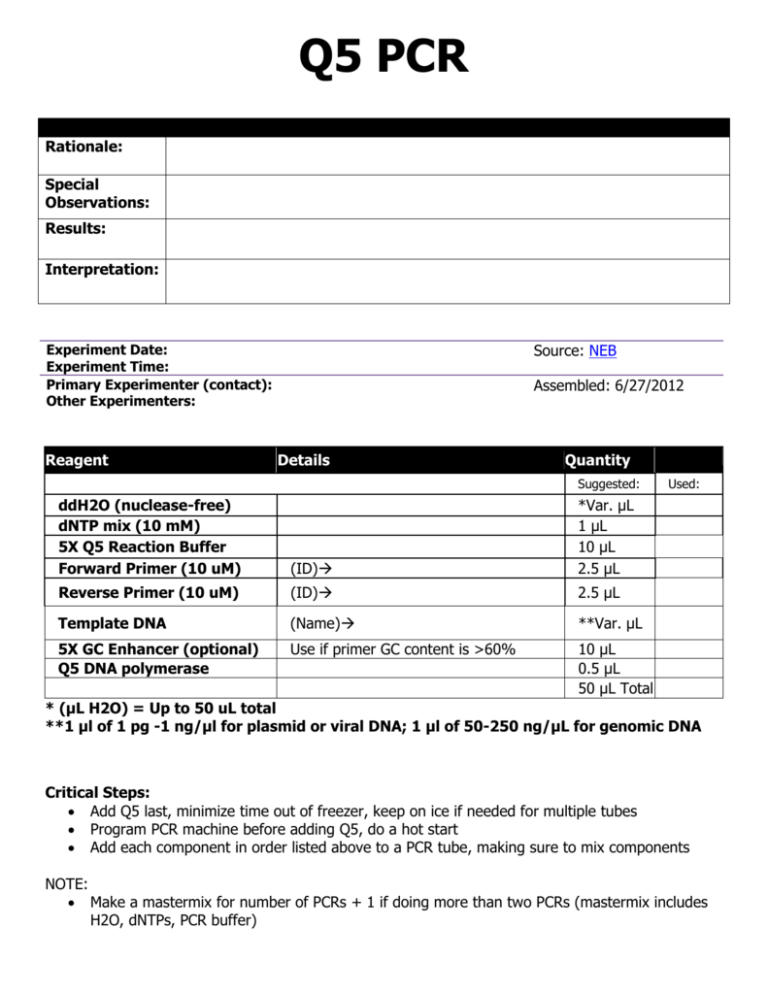




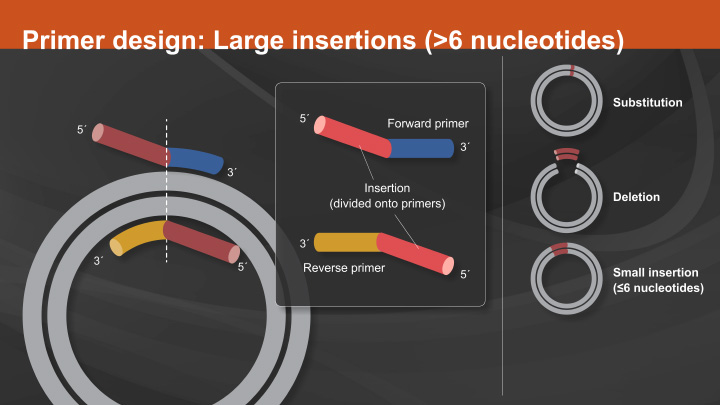









![PDF] Multiplex PCR using Q5® High-Fidelity DNA Polymerase | Semantic Scholar PDF] Multiplex PCR using Q5® High-Fidelity DNA Polymerase | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/637c0262c1bff0de8a5185963fb4c93d820d1bc6/1-Table1-1.png)
